 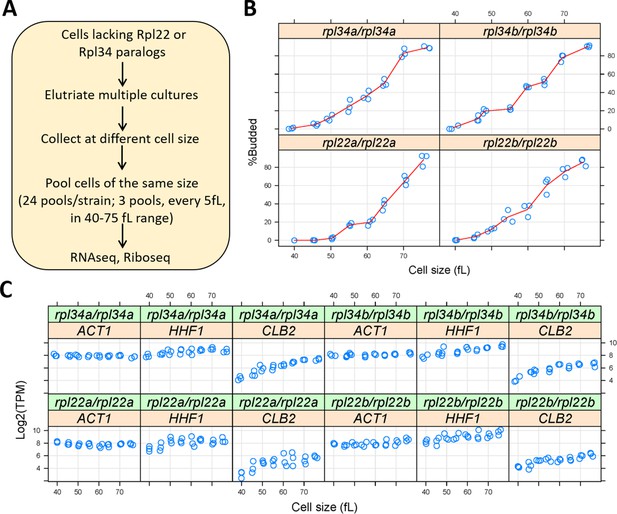
Observed differences are indicated with red upward arrows for segments that show relative deprotection while blue downward arrows indicate relative protection in the variant with respect to wild type. The level of protection is indicated by coloring each peptide at any given time-point based on the maximum percentage of deuterons incorporated according to the key on the right. The figure was generated by mapping all reliably-identified peptides onto Csk's wild type sequence then overlaying the resulting blocks with exactly matching peptides that have the same mass, charge and retention times in LC-MS analysis for both proteins. Regions of secondary structure motifs and domains identified from the crystal structure are shown above the sequence. Heatmap schematic of % Hydrogen-Deuterium exchange showing relative amide proton protection from D 2O solvent in full length variant and wild type Csk.Ĭolor-coded blocks show levels of deuteration for peptide probes in wild type Csk (top row) and the variant (bottom row) as indicated on the right. The reactions typically included 200 nM Csk, 20 µM kdSrc, and 50 µM ATP at 23☌. Wild type Csk's (circle) and the variant's (square) kinase activity was monitored in a ATP coupled radioactive assay in which a kinase dead substrate (kdSrc) is phosphorylated as a function of time. Structural alignment was performed in PyMol using PDB entries 2ETZ, 2SRC, and 3EAC for Itk, Src, and Csk, respectively.ĬD loop elongation in the SH2 domain results in reduced kinase activity of full length Csk towards its physiological substrate Src. Sequence alignment was performed using Clustal Omega through UniProt web server. Most notably, Csk possesses the shortest CD loop (turn) of the three modular domains. (B) Sequence and structural alignment of SH2 domains of homologous protein kinases: Itk (red), Src (blue), Csk (green) illustrating the stark differential loop lengths among these domains. (A) Structure and domain arrangement in full-length Csk (PDB ID 1K9A). Use the mobile_state and target_state argument to be explicit in such cases.Sequence and structural alignment of homologous SH2 domains. By defaults, all states (like in NMR structures or trajectories) are considered, this might yield a bad or suboptimal alignment for a single state.You can skirt this problem by making a temporary object and aligning your original to the copy. Else, PyMOL will answer with: ExecutiveAlign: invalid selections for alignment. The molecules you want to align need to be in two different objects.Number of aligned atoms before refinement.Number of aligned atoms after refinement.align ( string mobile, string target, float cutoff = 2.0, int cycles = 5, float gap =- 10.0, float extend =- 0.5, int max_gap = 50, string object = None, string matrix = 'BLOSUM62', int mobile_state = 0, int target_state = 0, int quiet = 1, int max_skip = 0, int transform = 1, int reset = 0 ) Note that the output prints "RMS" but it is in fact "RMSD" and the units are Angstroms.Ĭmd. The RMSD can also be captured with a python script, see the API paragraph below. The all-atom RMSD can be obtained by setting cycles=0 and thus not doing any outlier rejection. The RMSD of the aligned atoms (after outlier rejection!) is reported in the text output. can be saved to a clustalw sequence alignment file.graphical representation of aligned atom pairs as lines in the 3D viewer.cutoff = float: outlier rejection cutoff in RMS Īn alignment object can be created with the object= somename argument.target = string: atom selection of target object.mobile = string: atom selection of mobile object.[, matrix [, mobile_state [, target_state Usage align mobile, target [, cutoff [, cycles 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
